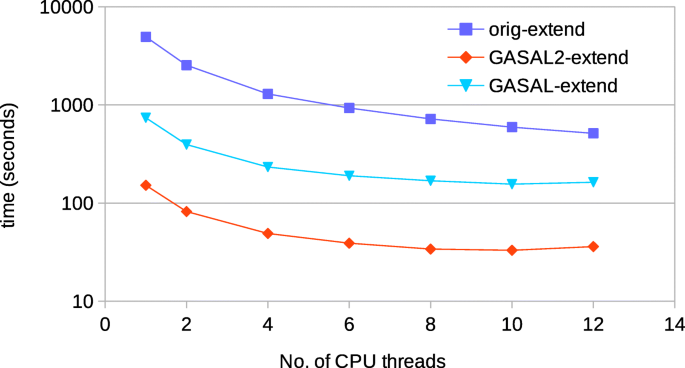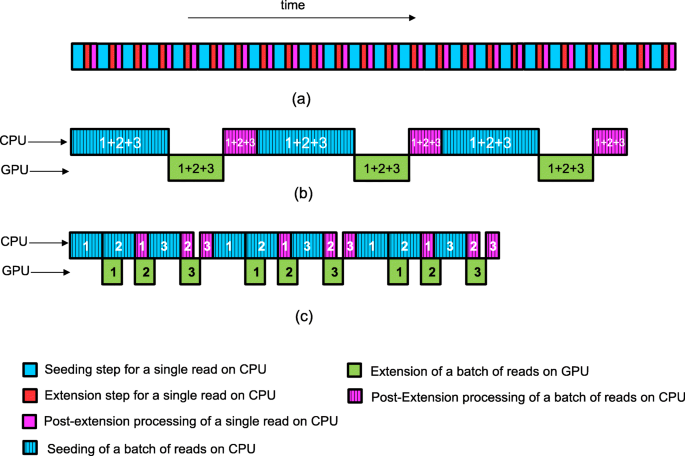
GASAL2: a GPU accelerated sequence alignment library for high-throughput NGS data | BMC Bioinformatics | Full Text
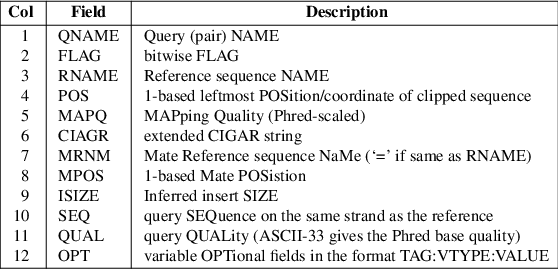
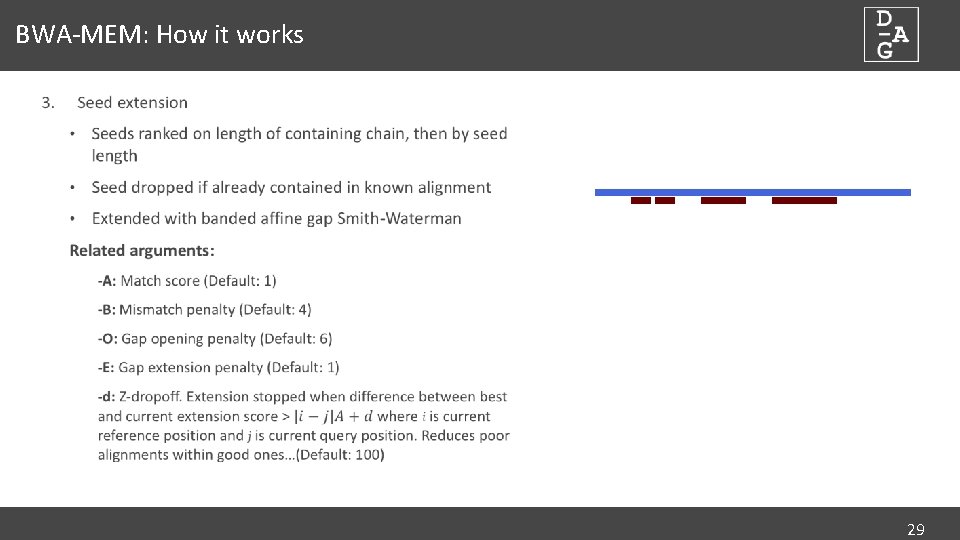
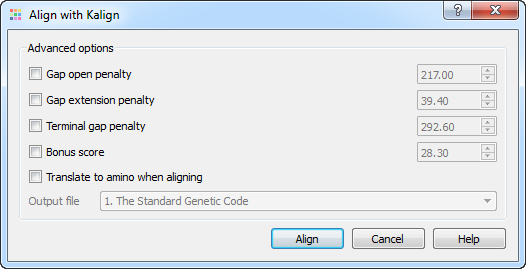
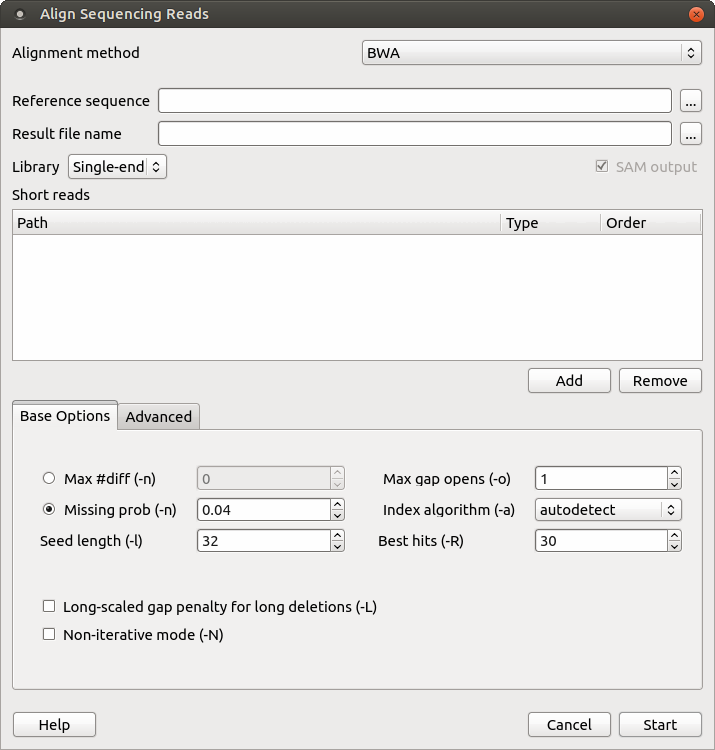
myExperiment - Workflows - cutadapt+BWA-MEM-mis1-i2,2-gapex2,2+filter_c5.0_c4.15+idxStats (Marc Ciosi) [Galaxy Workflow]

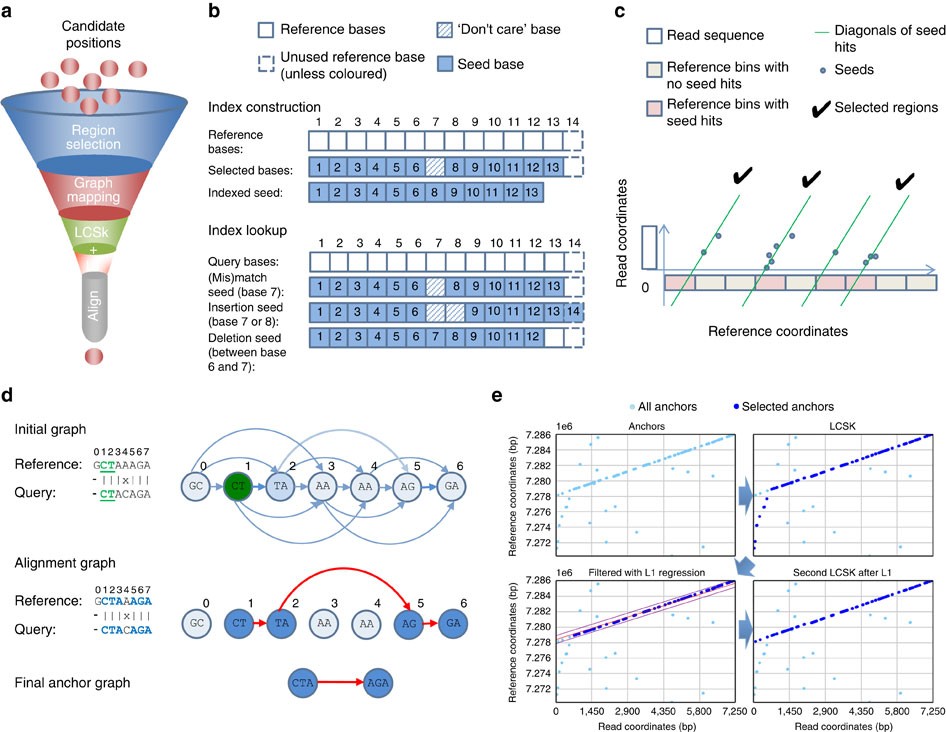
ParSNP Hash Pipeline to parse SNP data and output summary statistics across sliding windows. - ppt download